Research
Projects
Investigations across cell biology, genome architecture, and neuroimmunology. Click any project to expand.
Postdoctoral Work
MGB & Harvard Medical School · 2023–present
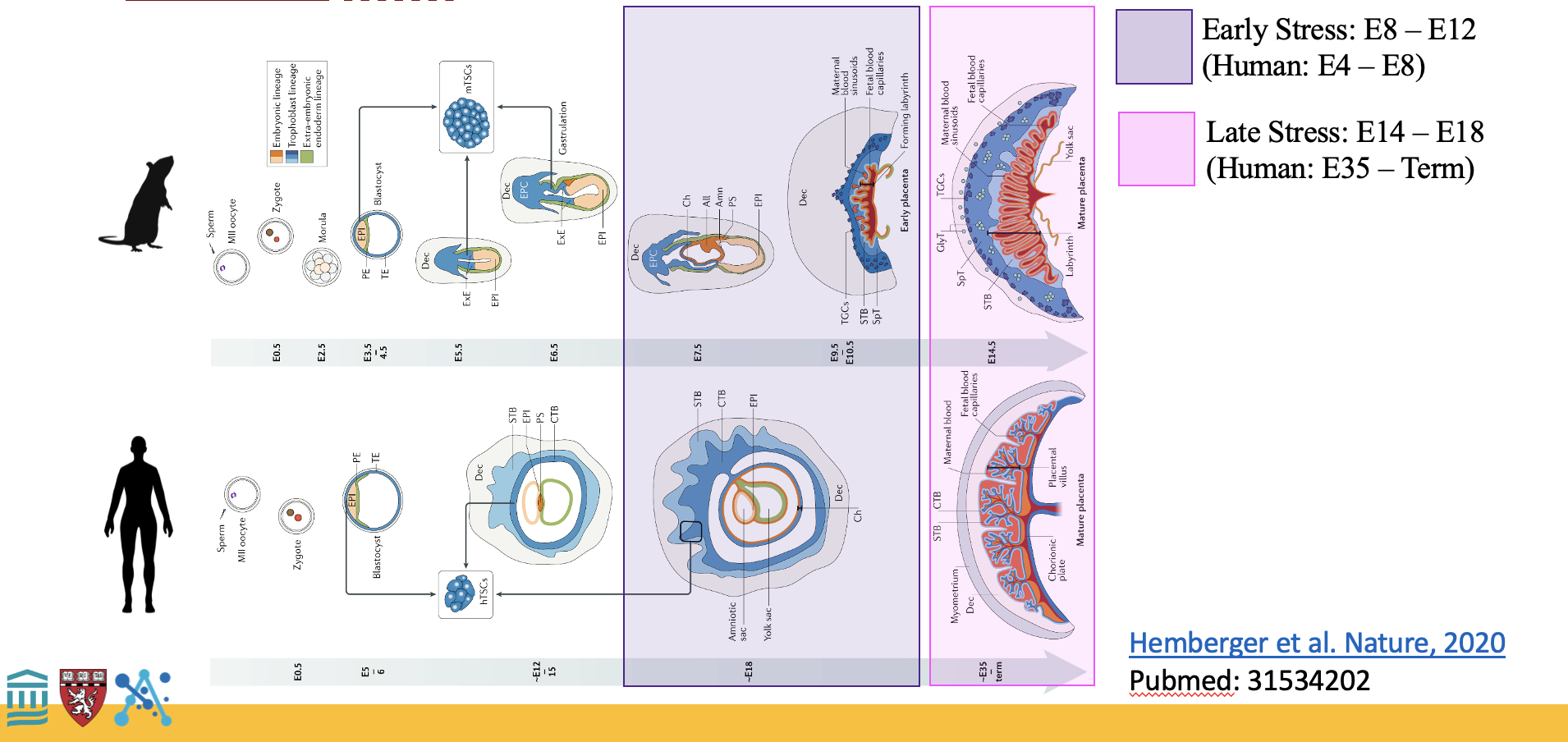
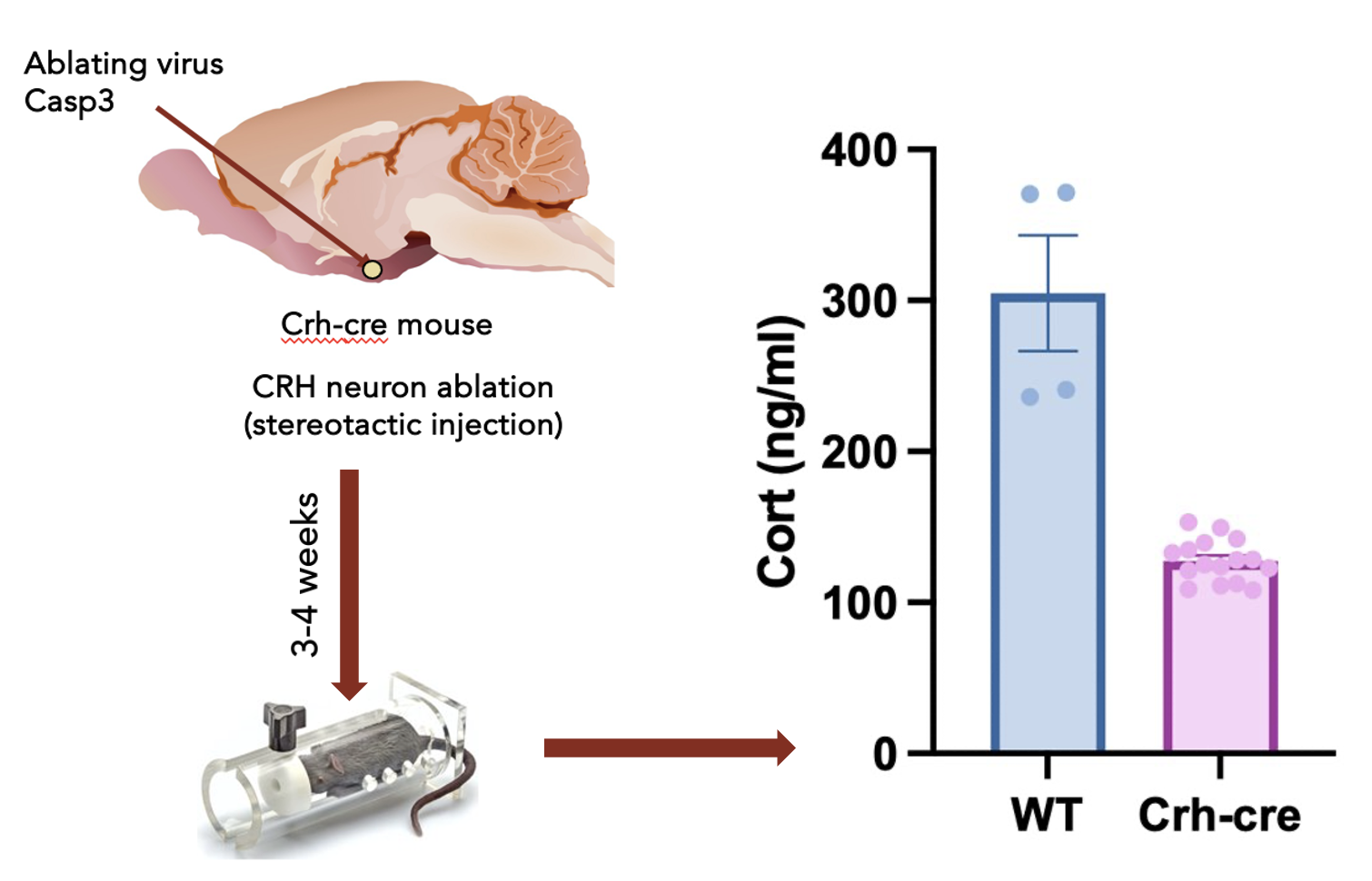
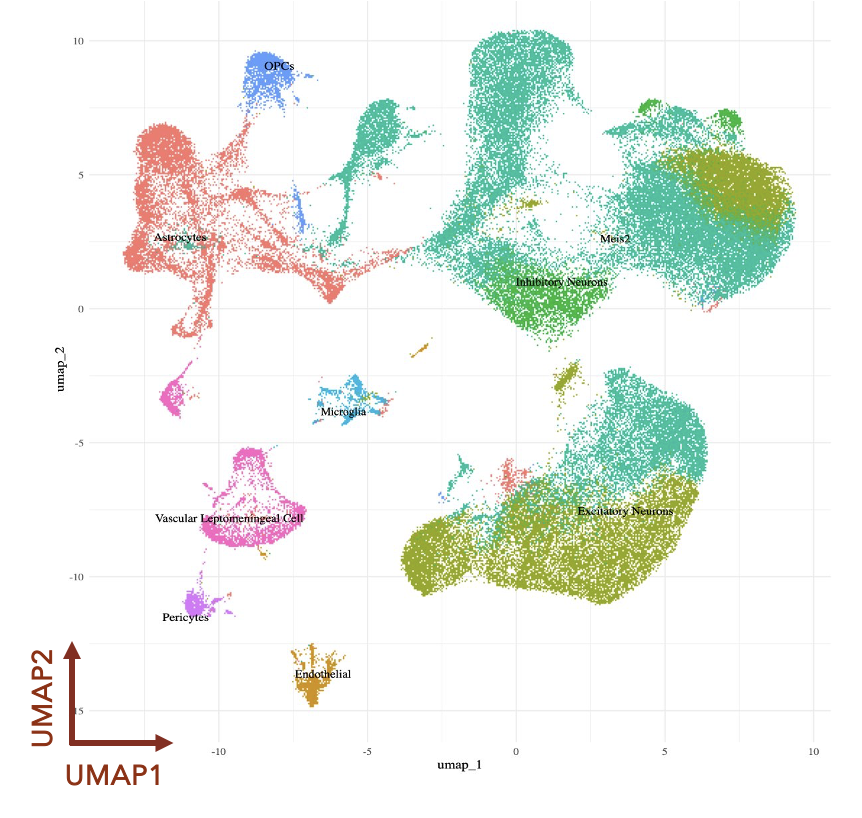
Prenatal Stress & Long-Term Health
Investigating how chronic prenatal stress reshapes immune function and brain development across a lifetime.
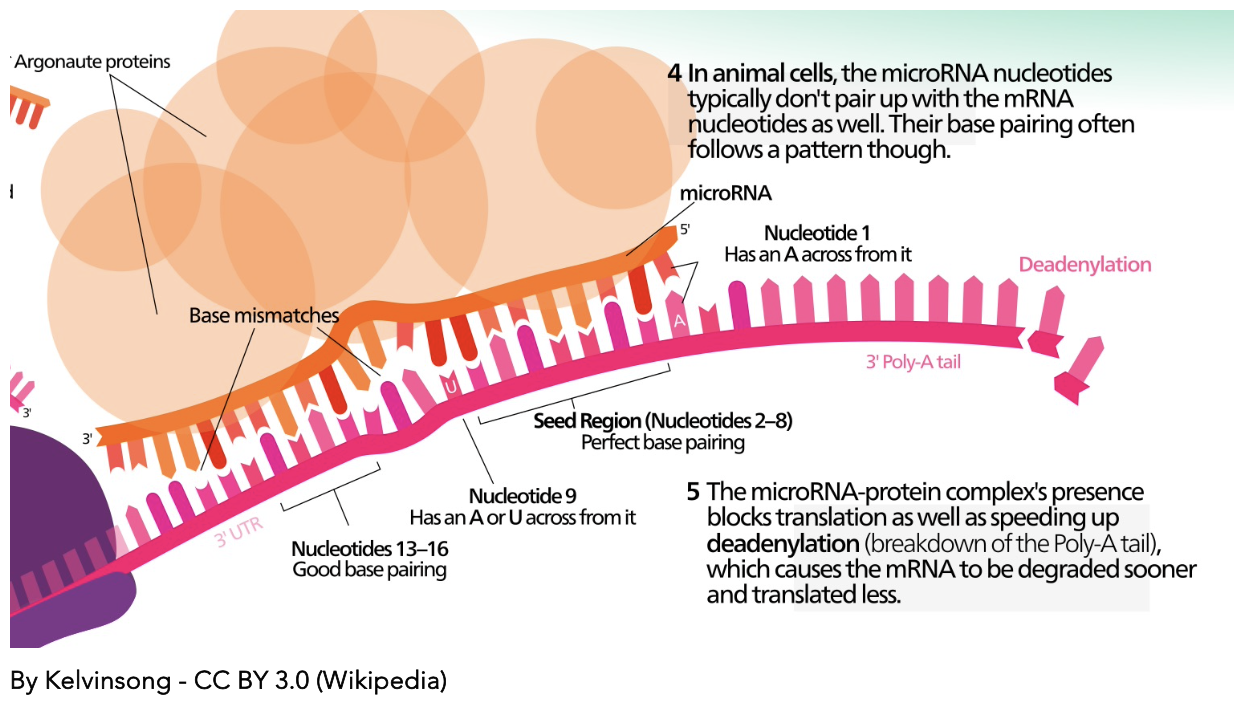
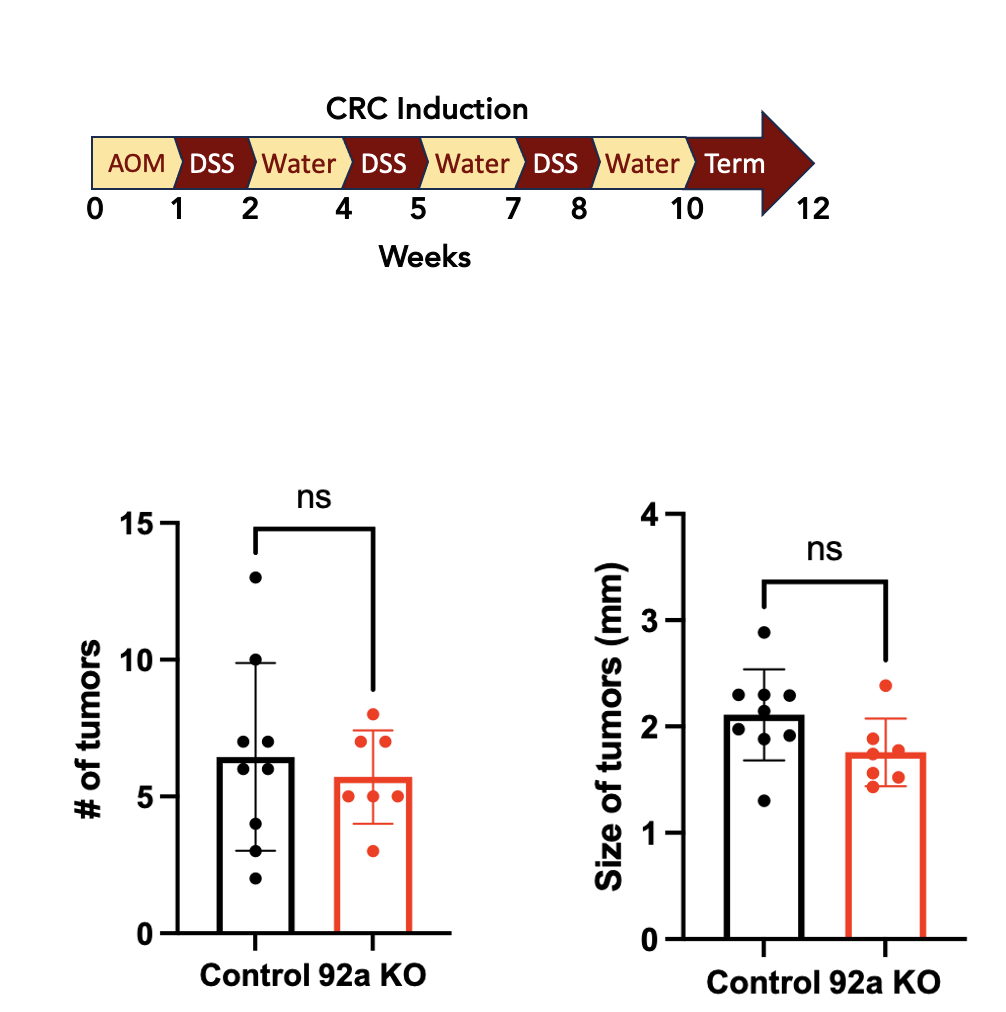
miRNAs in Cancers
Characterizing the roles of miR-92a and miR-21 in colorectal cancer, colitis, and Multiple Sclerosis.
PhD Work
Tata Institute of Fundamental Research · 2014–2019
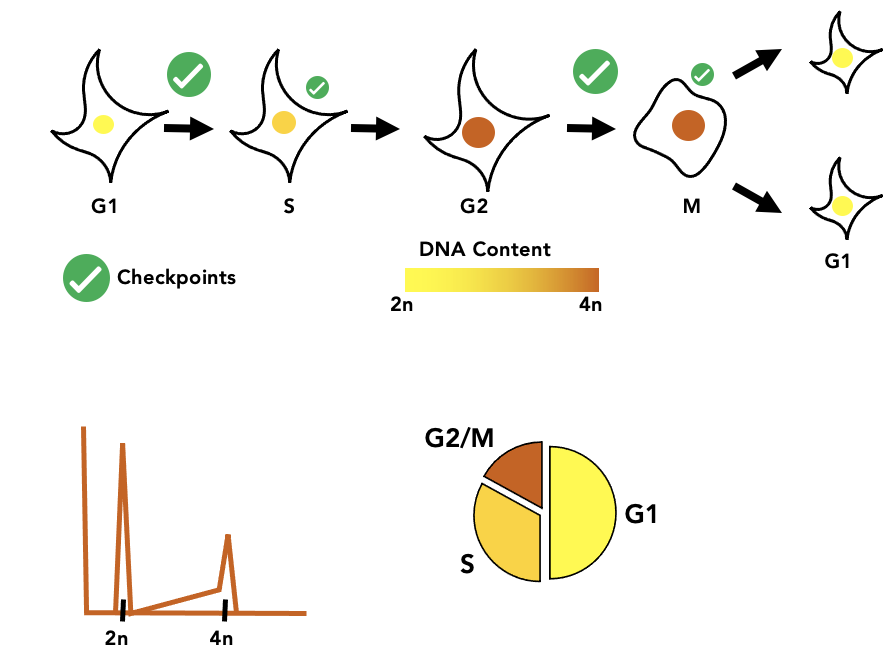
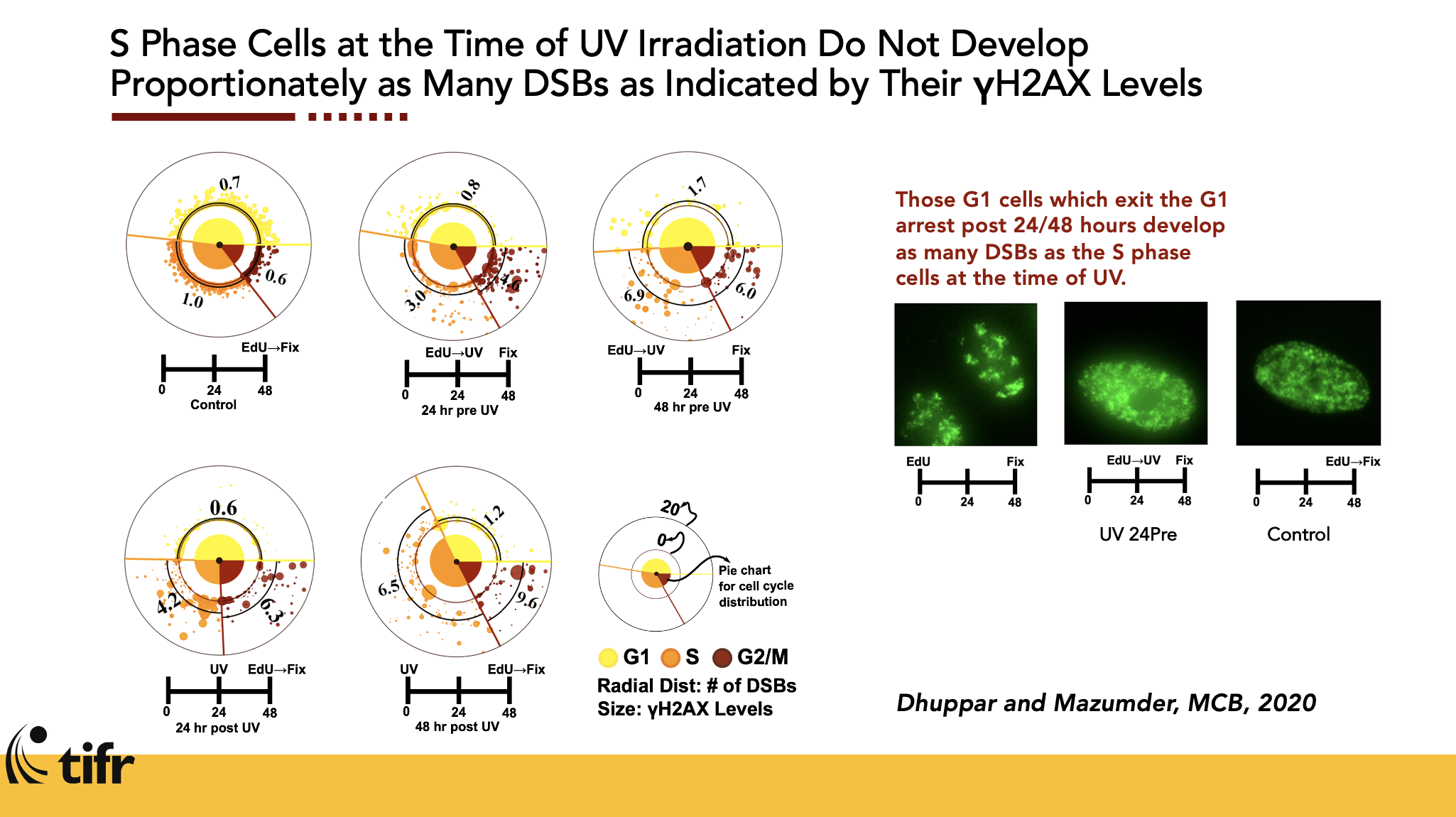
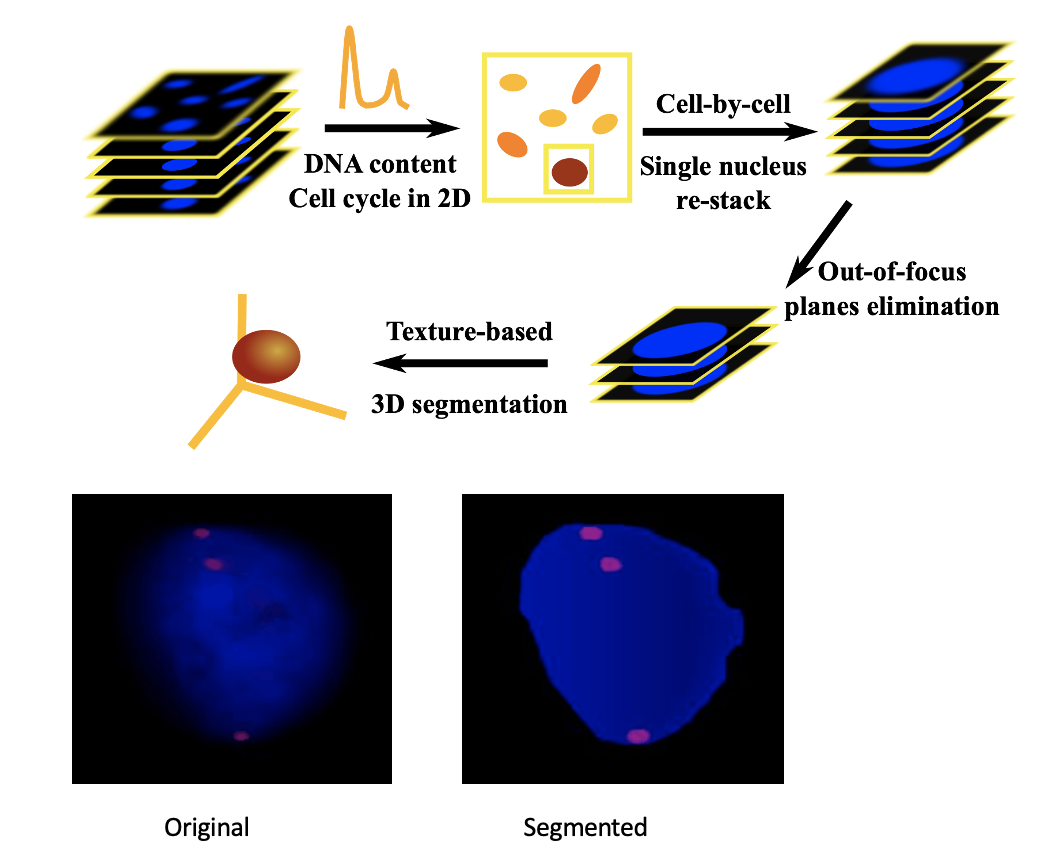
Cell Cycle Staging from Microscopy
Automated image analysis pipeline to stage cells in the cell cycle without chemical synchronization.
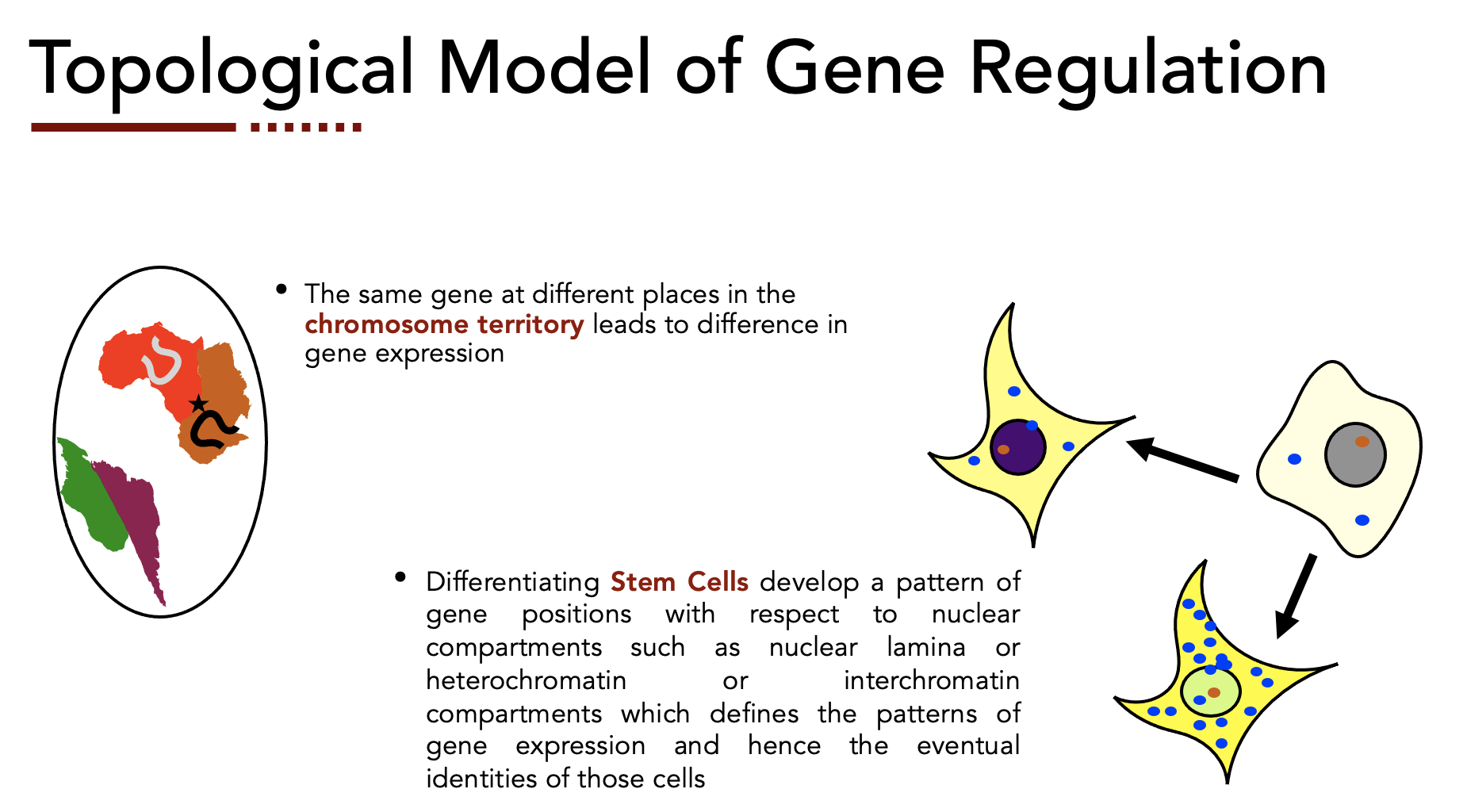

Genome Architecture & Gene Regulation
How CTCF-mediated three-dimensional genome organization governs monoallelic gene expression.